SciMed Education
DLS Applications for Nanoparticles, Proteins and Biotech Samples
In Summary
DLS characterises a wide range of nanomaterials—including liposomes, exosomes, polymers, micelles and protein aggregates—and provides insights into their size distributions, aggregation state and molecular weight.
Can DLS measure liposomes, lipid nanoparticles and exosomes?
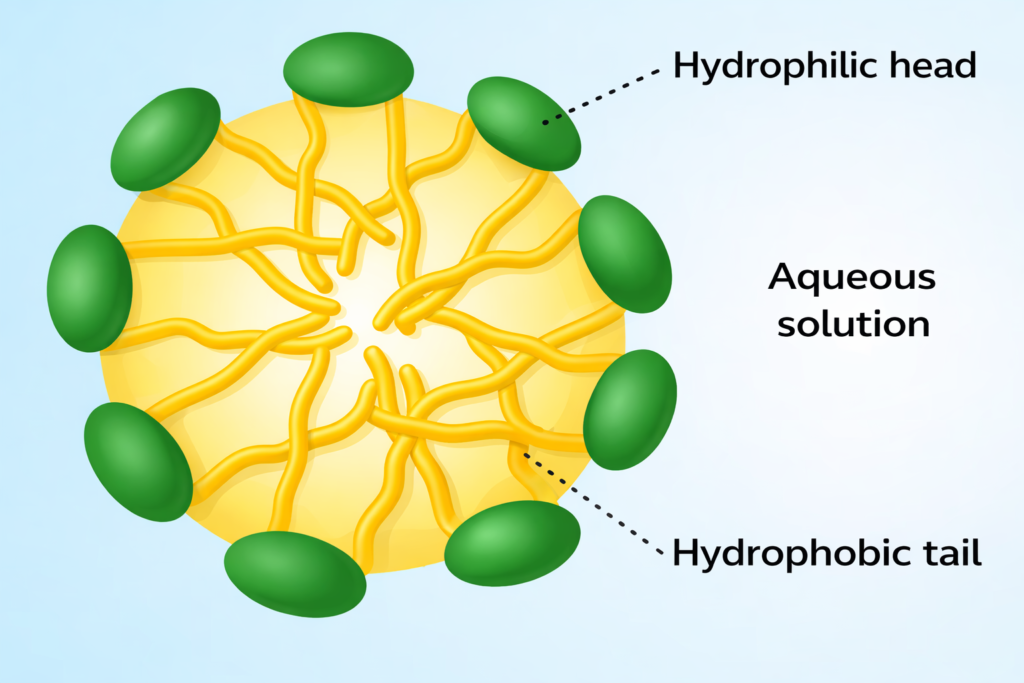
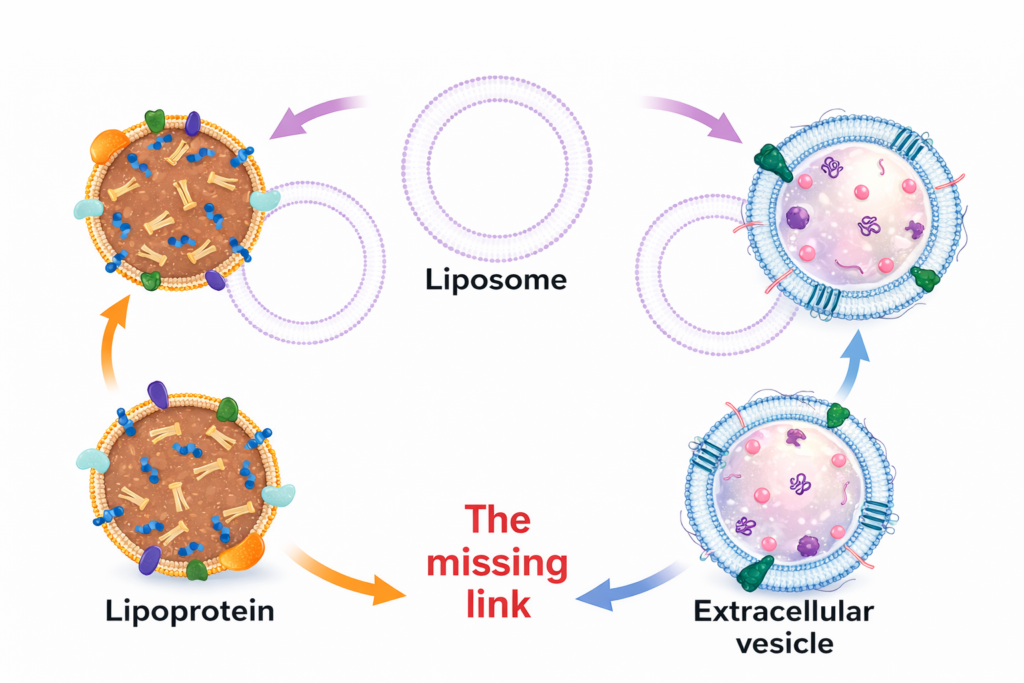
Liposomes and lipid nanoparticles, typically 80–250 nm in size, produce strong scattering and yield narrow distributions. Exosomes (40–160 nm) and extracellular vesicles fall well within the DLS range and are often monodisperse. Accurate size measurement requires careful sample preparation to remove protein or media contaminants.
How does DLS measure polymer nanoparticles and determine molecular weight?
DLS readily measures polymer nanoparticles and micelles down to a few nanometres. When combined with static light scattering, DLS data enable calculation of molecular weight and radius of gyration for polymers. Micelle sizes around 4.5 nm can be measured over wide concentration ranges—from as low as 0.5 % up to 30 %—using the DTC head.
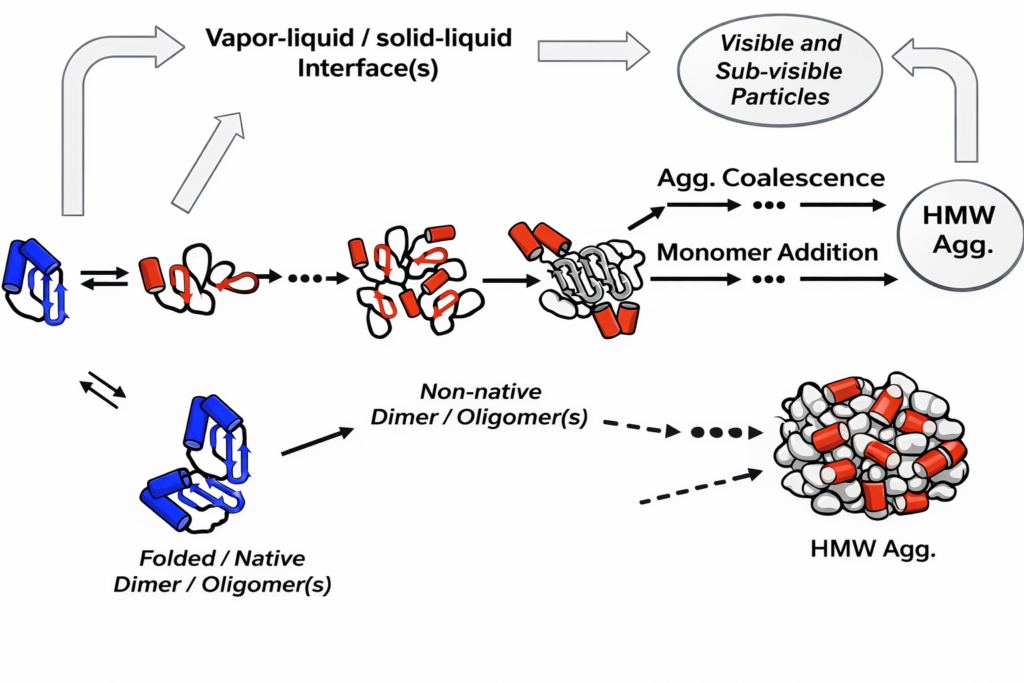
Can DLS detect protein aggregation and measure biological samples?
DLS is extremely sensitive to aggregation because scattering intensity scales with particle diameter to the sixth power. It can detect early‑stage aggregation in protein formulations.
Measurements in biological buffers such as phosphate‑buffered saline are possible if samples are transparent; the instrument needs solvent viscosity and refractive index to calculate accurate sizes.
How does DLS compare with nanoparticle tracking analysis (NTA) and can it measure zeta potential?
Standard DLS reports an ensemble‑average size, while NTA tracks individual particles and provides number‑weighted distributions. DLS covers a wider size range (~1 nm to 5 µm) and operates at higher concentrations; NTA is better for multi‑modal distributions and lower concentrations.
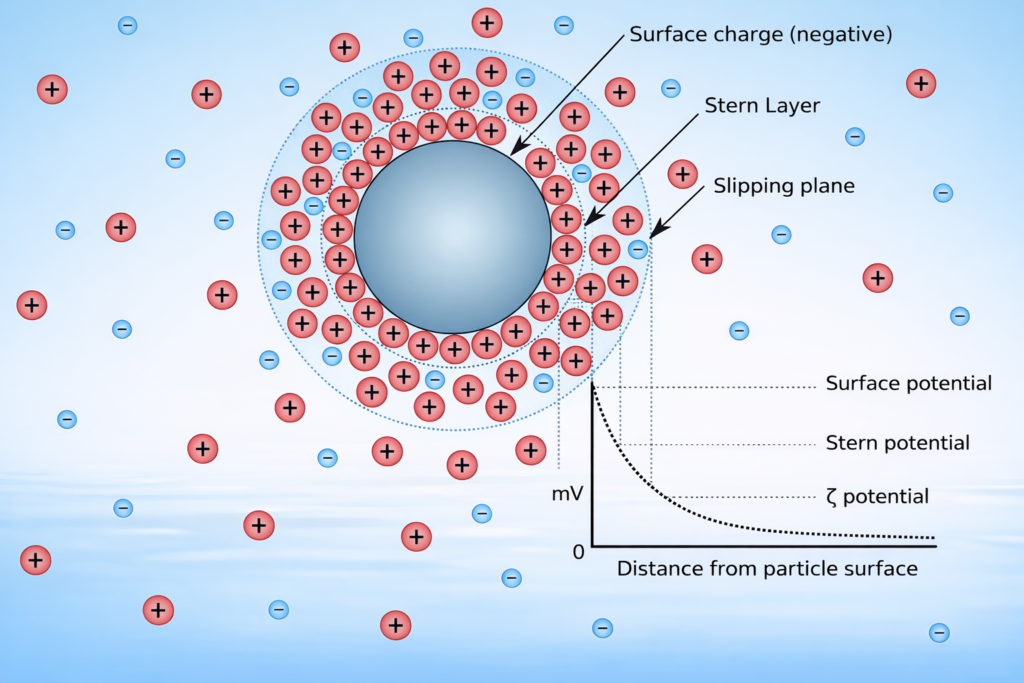
To measure zeta potential, DLS instruments can be paired with electrophoretic light scattering in a single instrument.
What about non‑spherical, anisotropic or fluorescent particles?
Traditional DLS assumes spherical particles, reporting an equivalent spherical diameter. Depolarised DLS (DDLS) measures depolarised light along perpendicular axes to estimate aspect ratio and size simultaneously.
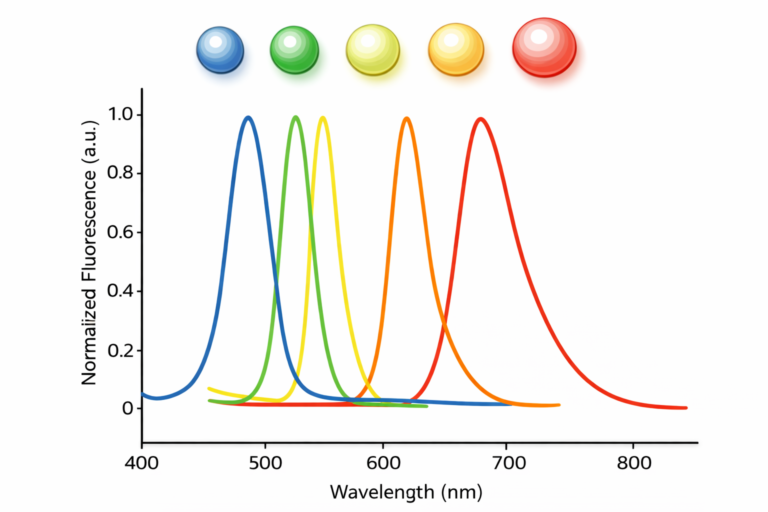
For fluorescent particles like quantum dots, longer‑wavelength lasers (e.g., 785 nm) combined with fluorescence filters enable accurate size measurement despite emission.
Can DLS measure complex biological matrices and quantum dots?
Yes. Quantum dots (1–20 nm) can be measured by choosing appropriate laser wavelengths and filters. Serum and blood samples require dilution, centrifugation and filtration to remove cells and aggregates; longer wavelengths reduce absorption and scattering. DLS can then quantify the size of lipoproteins and extracellular vesicles in these complex fluids.
What to do Next?
Whether you work in nanomedicine, polymer science or protein formulation, dynamic light scattering can accelerate your research by providing rapid size and aggregation information. Contact us to discuss measuring liposomes, exosomes, polymers, micelles or protein aggregates. Our VASCO Kin analyser supports these applications with interchangeable heads and combined DLS/SLS capabilities. Enquire now to learn more.
Page FAQ's
Poor preparation can introduce dust or aggregates that distort results.
By filtration or centrifugation before measurement.
It depends on the system, but both very low and very high concentrations can affect accuracy.
- Yes, because it changes solvent viscosity, which impacts calculated particle size.
Contact Us Today
We take great pleasure in assisting you and ensuring you get a prompt response to your questions
Live chat opening hours Mon – Fri 9:15 to 16:30 (UK Time)